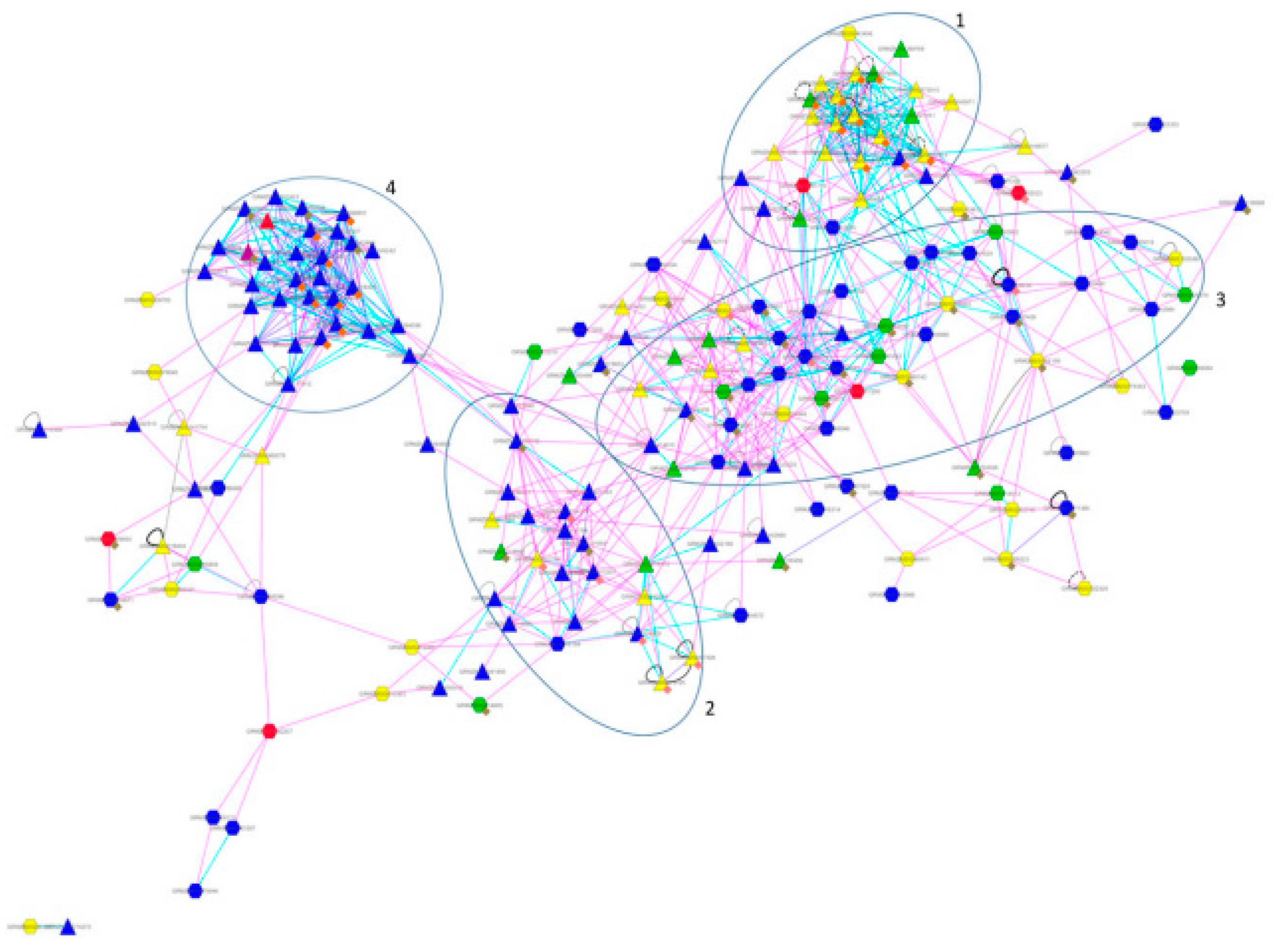 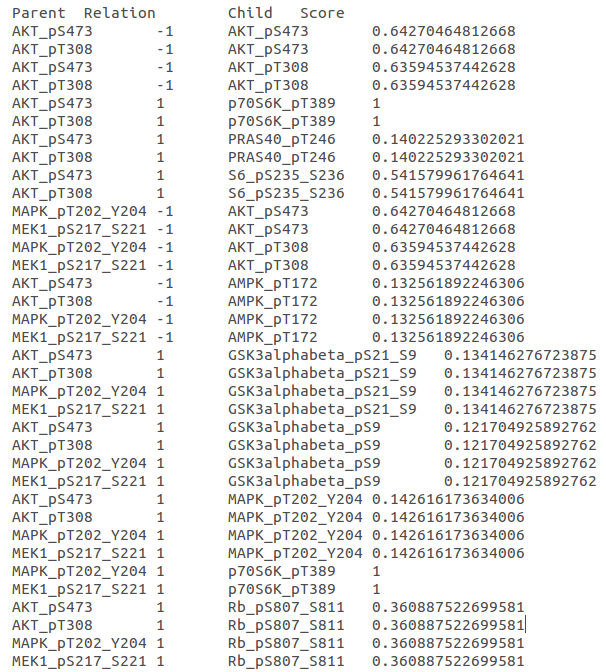
This study aimed to identify immune-associated biomarkers to complement the known predictive indicators and reveal the differential immune cell infiltration statuses between the HF and non-HF groups. discovered candidate time series differentially expressed genes (DEGs) associated with post-AMI HF, namely FADS2, LRRN3, GPR15, and AK5. revealed potential biomarkers for predicting post-AMI HF, including FMN1, JDP2, and RNASE1. However, the immune mechanisms of these genes in post-AMI HF have not been elucidated. īioinformatics approaches have identified several candidate genes for post-acute myocardial infarction (MI) (AMI) HF. Additionally, CIBERSORT enables the elucidation of the immune cell landscape in tumors. Cell-type identification by estimating relative subsets of RNA transcripts (CIBERSORT), a widely used algorithm for identifying immune cells, enables the assessment of relevant subpopulations of RNA transcripts and the quantification of cell components. Recently, bioinformatics approaches, a type of machine learning, have been used to identify biomarkers for the diagnosis and therapy guidance of HF. So we attempted to identify potential immune-related indicators for HF early diagnosis and therapy guidance. However, limited studies have identified the immune-associated biomarkers for predicting HF. Previous studies have demonstrated the role of immune response in HF pathogenesis. These findings indicate the potential correlation between myocardial diseases, cytokines, and immune cell populations. 
Additionally, the immune response in the impaired myocardium is characterized by the activation of the classical human immune response, which is similar to the response to autoimmunity and infection. T cells are reported to mediate the progression of pressure overload-induced HF. Under stress conditions, cardiomyocytes release proinflammatory cytokines, which trigger immune responses (including macrophage infiltration into the heart ) and mediate disease progression. Inflammation is involved in the pathogenesis of heart failure (HF). This study successfully identified two IRHGs that were significantly and differentially expressed in the HF group and could predict long-term HF, providing novel insights for future studies on HF and developing novel therapeutic targets for HF. Quantitative real-time polymerase chain reaction revealed that the Fos mRNA levels, but not the Cxcr5 mRNA levels, were downregulated in the mice of the HF group. Functional enrichment analysis revealed that the hub genes were enriched in immune system processes, including the interleukin-17 and nuclear factor-kappa B signaling pathways, which are involved in the pathogenesis of HF. The expression levels of CXCR5 and FOS and their ability to predict long-term HF were analyzed. This study identified two potential immune-related hub genes (IRHGs), namely CXCR5 and FOS, using bioinformatic approaches. However, limited studies have reported predictive immune-associated biomarkers for HF. Previous studies have reported the critical role of immune response in HF pathogenesis. Post-myocardial infarction heart failure (HF) is a major public health concern.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
